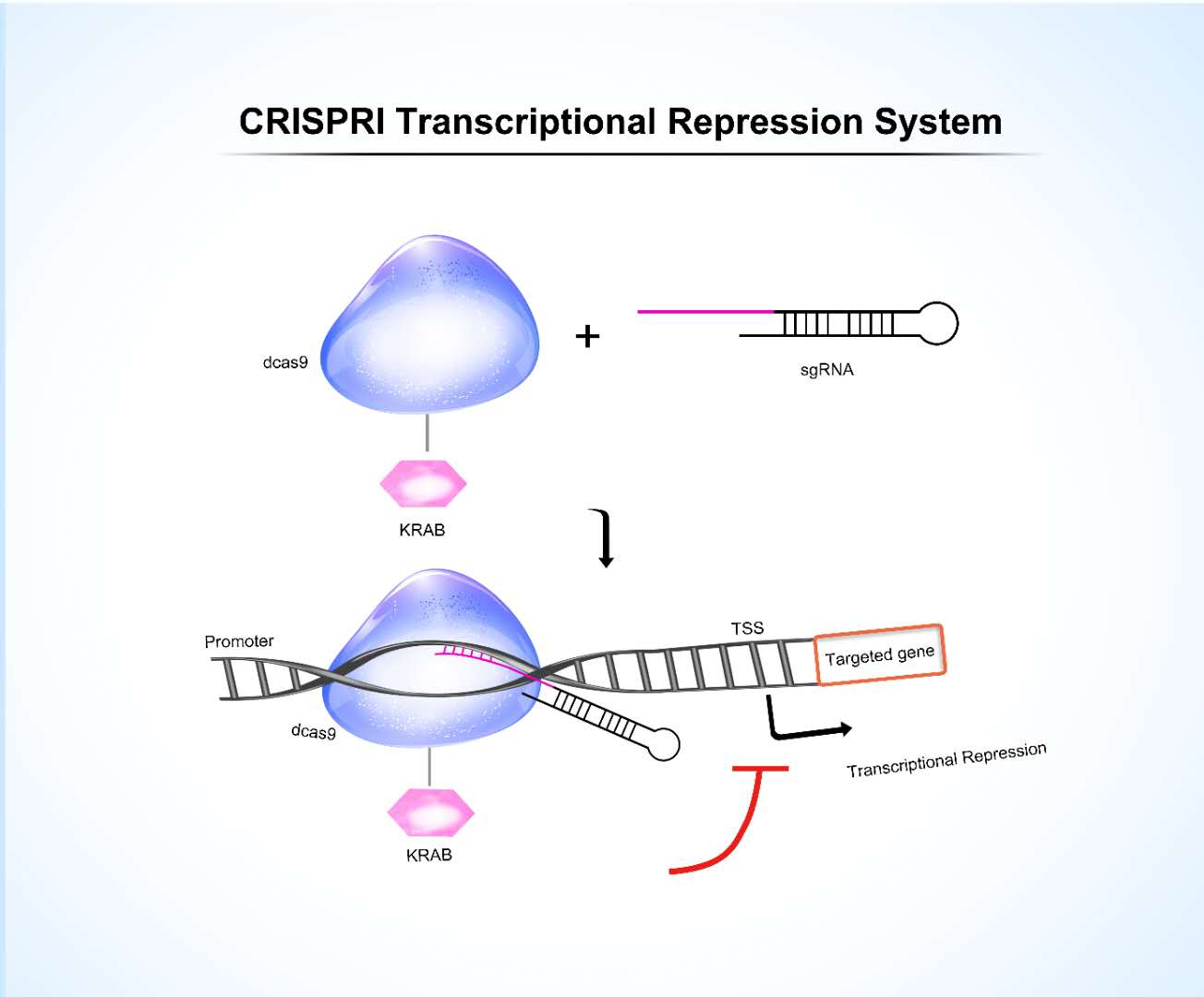
Repress gene expression, without cutting the DNA
CRISPR interference (CRISPRi) offers significant advantages over traditional gene knockdown methods like RNAi, including highly specific, reversible silencing of genes at the DNA level without altering the genome sequence. It provides high-efficiency repression, low off-target effects, and high-throughput functionality for studying essential genes, noncoding RNAs, and complex pathways.
GeneCopoeia’s CRISPRi products and services provide a complete, powerful solution to your epigenetic editing needs.
sgRNA clonesIntended for use with dCas9-KRAB |
sgRNA lentivirus particlesTransduction-ready particles, available in 3 purity levels |
Vector choices
| Vector | Promoter | CRISPR option | Selection Marker / Reporter Gene | Vector Type | Vector Map |
|---|---|---|---|---|---|
| pCRISPR-AB04 | U6 | CRISPRi dCas9-KRAB | Puro / mCherry | Lentiviral |
Technical overview
The KRAB system is the most classic CRISPRi tool, achieved by fusing the Krüppel-associated box (KRAB) repression domain to nuclease-dead dCas9. Guided by an sgRNA, dCas9-KRAB specifically binds near the transcription start site (TSS) of the target gene. The KRAB domain then recruits the KAP1 co-repressor protein, initiating a cascade of epigenetic modification events. KRAB-CRISPRi is particularly useful for loss-of-function screening of essential genes, studying non-coding RNA regulation, and phenotypic analyses that mimic partial drug inhibition effects.
The CRISPRi system comprises 2 components
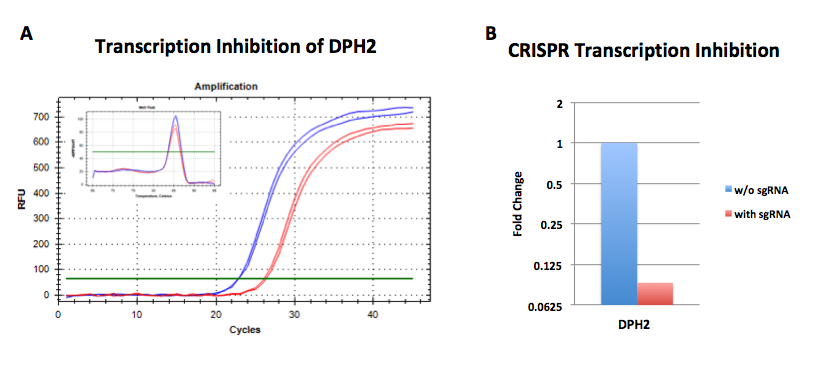 dCas9-KARB and sgRNA were co-overexpressed in HeLa cells. The transcriptional activity of DPH2 was assessed by qPCR. The amplification curves for cells transduced with sgRNA are shown in red, while those for cells without sgRNA transduction are shown in blue. A comparison of the transcriptional levels of the target gene with those of GAPDH demonstrated that the transcriptional levels of the target gene were significantly suppressed in HeLa cells.
dCas9-KARB and sgRNA were co-overexpressed in HeLa cells. The transcriptional activity of DPH2 was assessed by qPCR. The amplification curves for cells transduced with sgRNA are shown in red, while those for cells without sgRNA transduction are shown in blue. A comparison of the transcriptional levels of the target gene with those of GAPDH demonstrated that the transcriptional levels of the target gene were significantly suppressed in HeLa cells.
- dCas9-KRAB fusion protein. Guided by the sgRNA, the dCas9 protein binds to the target DNA. Its fused KRAB domain then recruits epigenetic enzymes, such as the histone deacetylases HDAC1/2 and the histone methyltransferases, by binding to co-repressor proteins, such as KAP1. This process represses the transcriptional level of the target gene.
- sgRNA
Application
 dCas9-KARB and sgRNA were co-overexpressed in HeLa cells. The transcriptional activity of DPH2 was assessed by qPCR. The amplification curves for cells transduced with sgRNA are shown in red, while those for cells without sgRNA transduction are shown in blue. A comparison of the transcriptional levels of the target gene with those of GAPDH demonstrated that the transcriptional levels of the target gene were significantly suppressed in HeLa cells.
dCas9-KARB and sgRNA were co-overexpressed in HeLa cells. The transcriptional activity of DPH2 was assessed by qPCR. The amplification curves for cells transduced with sgRNA are shown in red, while those for cells without sgRNA transduction are shown in blue. A comparison of the transcriptional levels of the target gene with those of GAPDH demonstrated that the transcriptional levels of the target gene were significantly suppressed in HeLa cells.