GeneCopoeia offers premade cancer model cell lines carrying diverse gene mutations commonly seen in tumor biomarkers, including EGFR, KRAS and BRAF in the MAPK pathway. The mutant cells are engineered based on the HCT116 cancer cell line, in which mutations are integrated either homozygously or heterozygously using CRISPR technology. These model cell lines can be used to enhance our understanding of cancer biology and the development of cancer therapies.
Order information
Choose from a list of premade cancer cell lines with your desired mutations. The mutant cell lines are provided along with the HCT116 wild type parental cell line.
Don’t see the mutant cell line you want? We also offer custom cell line service. Please contact us at inquiry@genecopoeia.com, or call 866-360-9531 or 301-762-0888 for a quote.
Product details
Cancer cell lines carrying hotspot mutations were engineered using CRISPR to introduce single-nucleotide mutations or in-frame indel mutations. The mutants are built with Cas9, sgRNA and a DNA donor fragment carrying the specific mutation and a puromycin marker; the marker is inserted into the nearby intron. Single clonal isolation were performed for sequence verification, and then expansion as shown below.
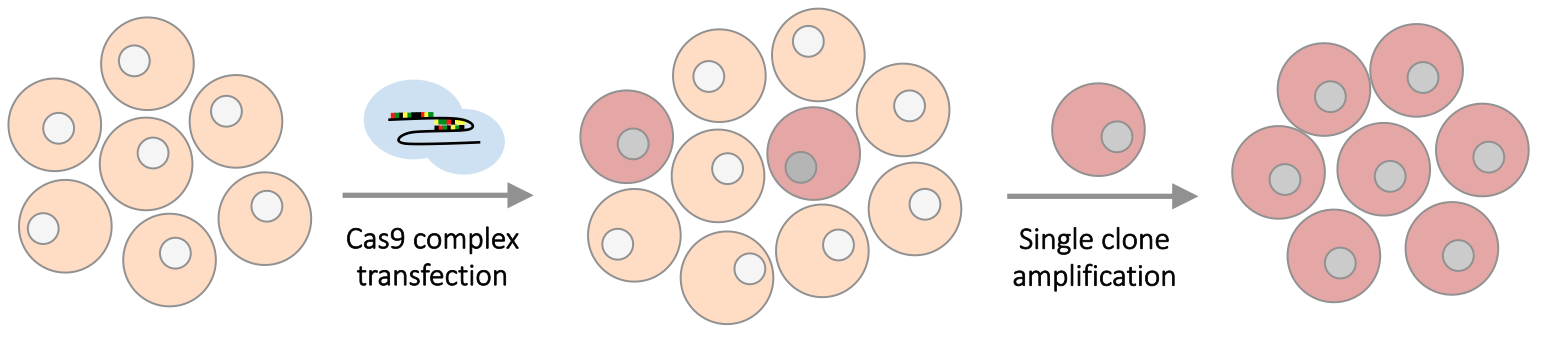 Figure 1. Cell line construction using CRISPR.
Figure 1. Cell line construction using CRISPR.
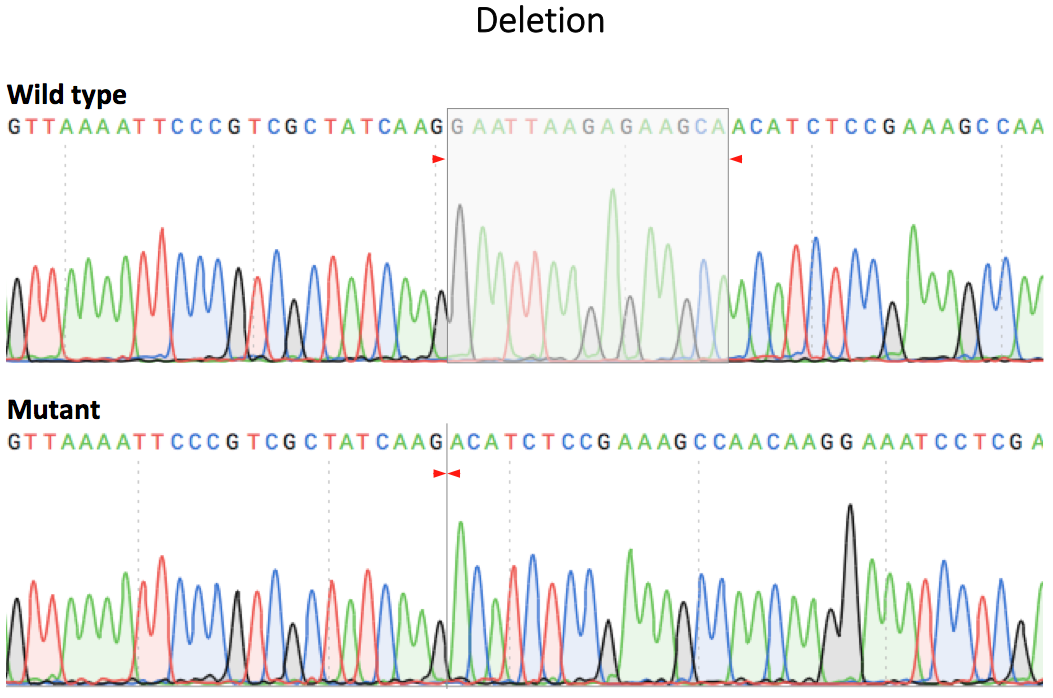
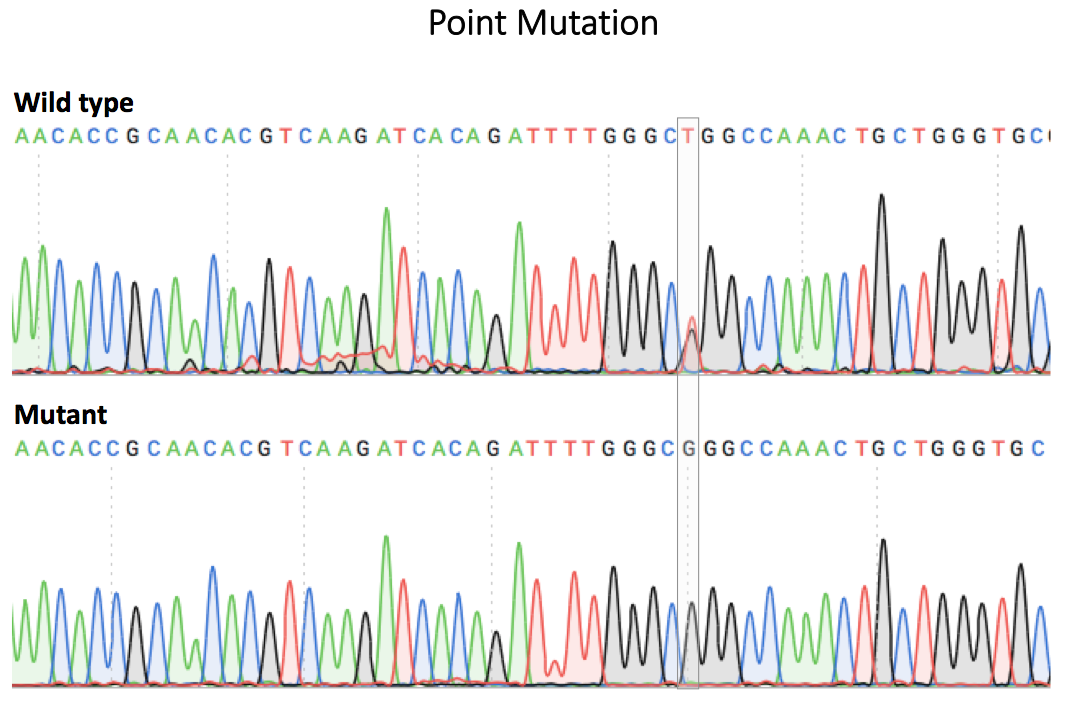 Figure 2. Sequence verification of a single cell clone using PCR and Sanger sequencing.
Figure 2. Sequence verification of a single cell clone using PCR and Sanger sequencing.
Comparison of the sequencing chromatograms of EGFR ΔE746-A750 cell line (left) and EGFR L858R cell line (right) to wild type. The deleted sequence in EGFR ΔE746-A750 and point mutation in EGFR L858R were marked in grey boxes in the chromatograms.
Technology overview
The mitogen-activated protein kinase (MAPK) pathway is critical for human cancer cell survival, proliferation and resistance to therapies. Genomic profiling has revealed hotspot mutations in the MAPK pathway components in tumors, such as EGFR, BRAF and KRAS genes. Mutations in these genes lead to increased protein amplification and alter the tumor microenviroment, therefore overactivate the downstream MAPK pathway. Drug therapies are developed with overactivated and oncogenic drivers of the MAPK pathway as therapeutic targets.
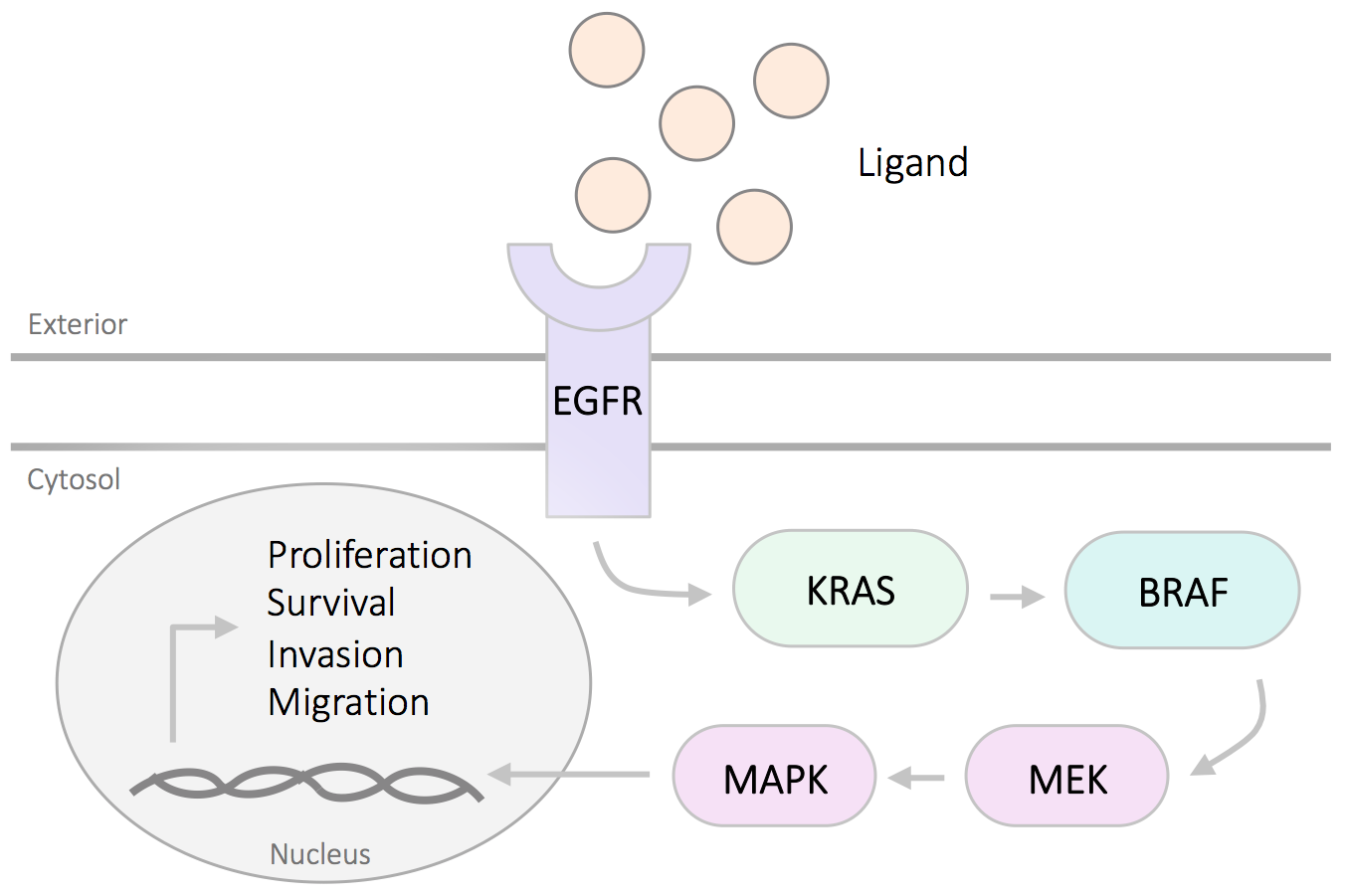
EGFR (epidermal growth factor receptor) is a cell surface tyrosine kinase receptor that binds to epidermal growth factor. Dysregulation of the EGFR contributes to a range of cancers and in various ways. EGFR has become an important therapeutic target for the treatment of NSCLC. The in-frame indel mutation ΔE746-A750 of exon 19 and point substitution L858R of exon 21 are the most common mutations in
EGFR, accounts for ~45% and 35% of all EGFR mutations respectively. Current FDA approved tyrosine kinase inhibitors (TKIs) for advanced lung cancers require the presence of these mutations. Inhibition of mutant EGFR in preclinical models through TKIs unsettles the intracellular signaling cascade, generating cell cycle arrest and apoptosis [1]. KRAS are small GTPases that mediate downstream growth factor receptor signaling, therefore are critical for cell proliferation survival and differentiation. Activating mutations in KRAS resulting in constitutive activation of the GTPase protein in the absence of growth factor signaling, resulting in a sustained proliferation signal within the cell. Approximately 36-40% of patients with colorectal cancer have tumor associated KRAS mutations. The KRAS G13C mutation is strongly associated with colorectal cancer. Studies have shown that KRAS mutations are linked to poor response to anti-EGFR antibody therapy in colorectal cancer [2]. Therefore, it is recommended to test for KRAS mutations before applying EGFR-inhibiting drugs to colorectal cancer patients. BRAF belongs to the serine/threonine-protein family of growth signal transduction kinases. BRAF mutations can be found in some types of cancer, including melanoma and colorectal cancer. About 50% of melanomas harbors activating BRAFmutations, with over 90% of which V600E. The BRAF V600E mutation deregulates activation of the downstream MEK/ERK effectors, evades senescence, apoptosis and immune response, and drives the proliferation of cancer cells [3]. Kinase inhibitors targeting BRAF V600E mutation have gained FDA approval for the treatment of malignant melanoma and NSCLC.
References
- 1. Costa DB. Kinase inhibitor-responsive genotypes in EGFR mutated lung adenocarcinomas: moving past common point mutations or indels into uncommon kinase domain duplications and rearrangements. Transl Lung Cancer Res. 2016 Jun;5(3):331-7.
- 2. Lièvre A, Bachet JB, Le Corre D, Boige V, Landi B, Emile JF, Côté JF, Tomasic G, Penna C, Ducreux M, Rougier P, Penault-Llorca F, Laurent-Puig P. KRAS mutation status is predictive of response to cetuximab therapy in colorectal cancer. Cancer Res. 2006 Apr 15;66(8):3992-5.
- 3. Maurer G, Tarkowski B, Baccarini M. Raf kinases in cancer-roles and therapeutic opportunities. Oncogene. 2011 Aug 11;30(32):3477-88.




