Introduction
The IRES element is exactly what’s needed for certain experiments when a single-gene vector falls short. IRES enables the coordinated co-expression of two genes with the same vector. For example, you can monitor the delivery of one gene by using a second gene with a fluorescent tag or express a protein of interest and simultaneously biotinylate it with the same vector. An IRES element is especially useful for gene delivery when working with stem and primary cells and when the following tasks are pursued:
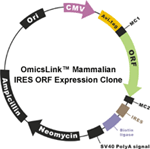
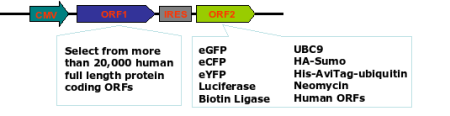 More than 20,000 human and 15,000 mouse genes are available in CMV promoter driven bicistronic (IRES) expression vectors.
More than 20,000 human and 15,000 mouse genes are available in CMV promoter driven bicistronic (IRES) expression vectors.
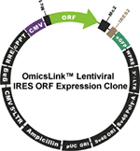
Figure1. OmicsLink™ ORF cDNA expression clones with IRES element in mammalian and lentiviral expression vector systems.
- Monitor gene delivery efficiency
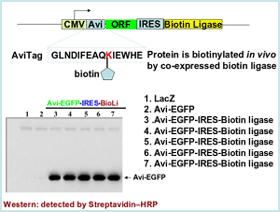
- In vivo biotinylation
- Stable transfection

Technology
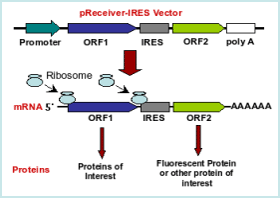
The IRES technology serves a dual-purpose. It allows the coordinated and efficient expression of two genes using the same promoter in a single vector (see figures 1 and 2). Virtually any combination of genes is possible. GeneCopoeia has already developed many constructs.Combined with OmicsLink expression-ready clones
For a variety of applications, GeneCopoeia has incorporated the IRES element into several mammalian and lentiviral vectors using a wide choice of reporter genes including the following:- eGFP
- mCherry
- SUMO
- biotin ligase
- neomycin
- luciferase
 More than 20,000 human and 15,000 mouse genes are available in CMV promoter driven bicistronic (IRES) expression vectors.
More than 20,000 human and 15,000 mouse genes are available in CMV promoter driven bicistronic (IRES) expression vectors.
Advantages
- Minimal effect on biological activities of target proteins when co-expressed with reporter/ assaying genes.
- Efficient co-transfection due to coordinated co-expression of two genes from a single expression vector.