|
Ed Davis, Ph.D. |
Biomedical researchers are enjoying a Renaissance in functional genomics, which aims to use a wealth of DNA sequence information—most notably, the complete sequence of the human genome—to determine the natural roles of the genes encoded by the genome. As a result, biochemical networks and pathways will be better understood, with the hope of leading to improved disease treatments.
A major approach of functional genomics is to ablate gene function, by either “knockdown” (reduction) or “knockout” (complete elimination). Since 2012, researchers have turned increasingly to CRISPR (clustered, regularly interspaced, short palindromic repeats) for functional genomics studies. CRISPR’s simple RNA-guided mechanism provides a quick, convenient, and relatively low-cost method for many applications, from gene knockout, in-frame fusion tagging, mutagenesis, and transgene knockin. Several groups recently adapted CRISPR for high-throughput knockout applications, by developing large-scale CRISPR sgRNA libraries. GeneCopoeia recently launched a number of smaller, pathway- and gene group-focused CRISPR sgRNA libraries, which offer several key advantages over the whole-genome libraries. In this Technical Note, we discuss the merits and applications for CRISPR sgRNA libraries, how to use CRISPR sgRNA libraries, the advantages of using small, pathway- and gene group-focused libraries, and how GeneCopoeia can help you with your high-throughput CRISPR knockout studies.
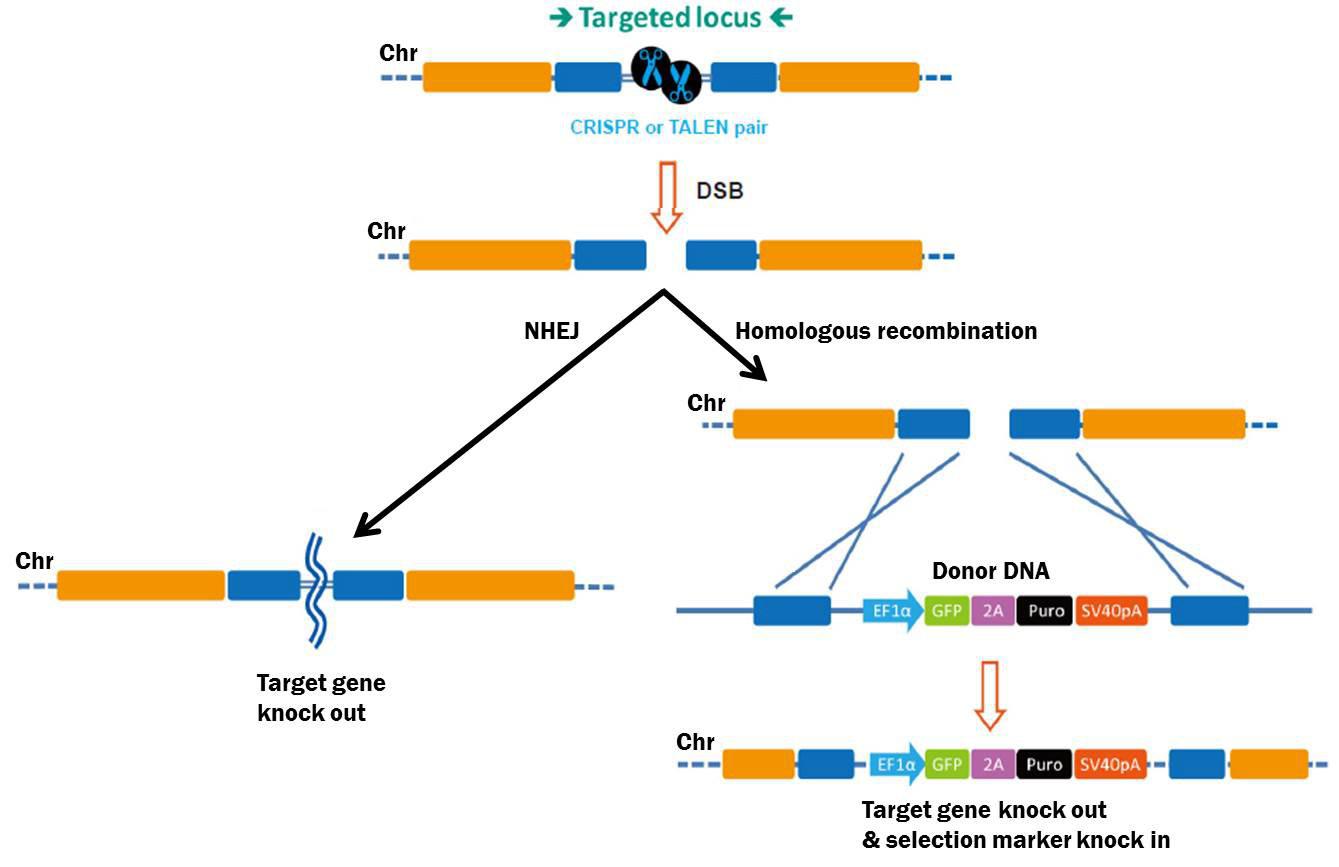
Figure 1. Pathways for repair of DSBs induced by genome editing tools. Left: NHEJ. Right: HR in the presence of a donor construct.
What are CRISPR sgRNA libraries?
CRISPR sgRNA libraries are collections of hundreds-to-thousands of plasmids, each expressing a unique chromosome target-specific single guide RNA (sgRNA), which, in the presence of Streptococcus pyogenes Cas9 causes double-strand breaks (DSBs) in the chromosome (Figure 1). DSBs are repaired either by homologous recombination (HR) in the presence of a homologous donor construct, or nonhomologous end joining (NHEJ), which leads to knockout mutations.
CRISPR sgRNA libraries enable researchers to knock out many genes simultaneously in mammalian cells, opening the door to many applications, such as drug target identification, drug target validation, phenotypic changes, and reporter assays. Previously, these applications were achieved using RNA interference (RNAi). For example, Sethi, et al., (2012) used an shRNA library directed against 6,000 “druggable” genes to identify potential targets in ovarian cancer cells. Cooper and Brockdorff (2013) used an shRNA library to identify factors that activated a Gata6-neo promoter reporter. Finally, Zhang, et al, (2014) used fluorescence activated cell sorting (FACS) to isolate cells expressing the cell surface marker PSA-NCAM, which is expressed in human embryonic stem cells (hESCs).
While shRNA libraries are useful for high-throughput loss-of-function screens, RNAi has a number of disadvantages compared with CRISPR: 1) RNAi causes a knockdown of the level of gene expression. Therefore, false negatives, resulting from residual gene expression, can be missed; 2) RNAi acts only on cytoplasmic RNA, and so cannot silence nuclear RNAs like long non-coding RNAs. Conversely, CRISPR makes permanent changes to the genetic code. Thus, CRISPR can make a complete knockout of all alleles of a gene, whether their transcription products are localized to the nucleus or cytoplasm.
How are CRISPR sgRNA libraries used?
Using sgRNA libraries is straightforward. However, it is important to have a “readout”, (observable phenotype or assay) in order for the libraries to be useful (Figure 2). Briefly, sgRNA libraries are generally used as pools of lentiviral particles, although the plasmids can also transfect cells as DNA. Our library clones contain sgRNA target sites, but not Cas9, which is required. So you will need to either co-transfect /co-transduce your cells with sgRNAs and Cas9, or transfect/transduce a cell line already stably expressing Cas9. For lentiviral transduction, we recommend first establishing a cell line stably expressing Cas9; the large size of Cas9 limits the viral titer, so transduction is more efficient with the sgRNA-expressing lentiviruses alone. In any case, cells are transfected with DNA or infected with the lentiviral pools, followed by screening for the response of interest. Examples of some screening readouts include:
- Cell death. Cells carrying lethal knockouts will be underrepresented in a pool of surviving cells.
- Drug resistance: Cells resistant to a drug of interest will be overrepresented in the pool after applying drug treatment.
- Reporter assay: Cells that lose the ability to express a fluorescent reporter (e.g. GFP) will be underrepresented in the population after FACS.
After the readout method or assay is applied, DNA sequencing is used to identify which sgRNAs are either under- or over-represented in the pools. For a very large, whole-genome scale library, it is necessary to use next-generation sequencing to identify these sgRNAs targeting candidate genes.
Development of CRISPR sgRNA libraries
In light of the advantages of CRISPR over RNAi for eliminating gene function, a few researchers have produced large CRISPR sgRNA libraries. In one study, Wang, et al. (2014) constructed pools of sgRNA-expressing lentivirus particles targeting 7,300 genes, which were used to infect a haploid cell line that was stably expressing Cas9 nuclease. In screening the transduced line for cells resistant to 6-thioguanosine, these researchers observed enrichment for cells carrying sgRNAs targeting genes in the mismatch repair pathway, but not in other DNA repair pathways. Similarly, Feng Zhang’s group at MIT used a lentiviral sgRNA library targeting approximately 18,000 genes to identify melanoma and stem cells that were either sensitive or resistant to vemurafenib (Shalem, et al., 2014). The Zhang lab has since followed up on that initial study by producing second-generation libraries that contain several improvements and target a larger number of genes in human and mouse. (Sanjana, et al. 2014).
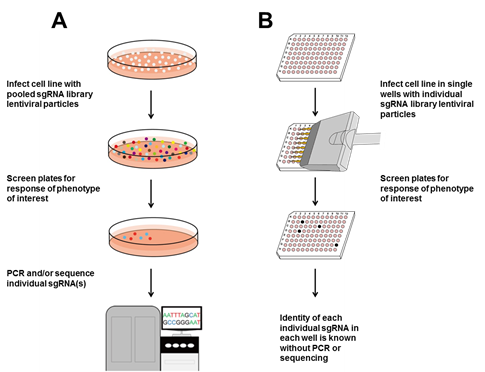
Figure 2. Basic workflow for using CRISPR sgRNA libraries. A. Pooled screen. Cells are infected with each sgRNA library pool. Infected pools are screened for the desired readout or phenotype. Pooled cells are subjected to Sanger sequencing for individual sgRNAs, or deep sequencing to look for over- or under-representation of individual sgRNAs. B. The GeneCopoeia GeneHero™ CRISPR sgRNA libraries are available either as pools or individual sgRNAs. If desired, cells can be infected with lentiviruses expressing individual sgRNAs. Plates or wells are screened for the readout or phenotype of interest. Individual sgRNAs corresponding to the phenotype of interest are already known without sequencing.
GeneCopoeia GeneHero™ CRISPR sgRNA libraries
Recently, GeneCopoeia launched a series of pre-made CRISPR sgRNA libraries, which are targeted to smaller gene groups and pathways (Table 1).
Table 1. GeneCopoeia GeneHero™ pre-made human CRISPR sgRNA libraries. Each library can be ordered as pools of either bacterial stock, transfection-ready DNA, or lentiviral particles from our website.
Each sgRNA is individually constructed, sequence-verified, and cultured in E. coli before pooling, in contrast to other CRISPR sgRNA libraries, which are constructed from the beginning as pools. GeneCopoeia’s CRISPR sgRNA libraries have several specific advantages over other sgRNA libraries, such as:
- Individual construction of each sgRNA makes it possible to purchase the libraries either as pools or as individual clones.
- Sequence verification provides high quality of each sgRNA.
- Individual construction and culturing of each sgRNA ensures the best representation in pools.
- Small library sizes reduce the amount of work and time spent on analyzing your results. You can screen the library for candidates using either Sanger or next-generation sequencing.
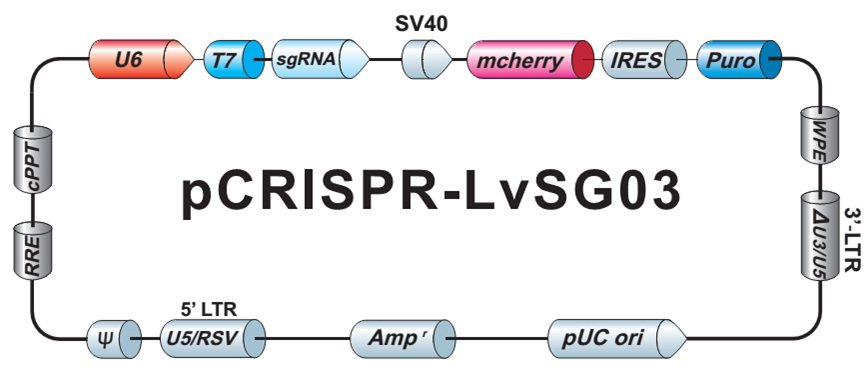
Our library sgRNAs are cloned into a very versatile vector backbone (Figure 3), which can be used for either direct transfection of cells, packaging into lentiviral particles, or in vitro transcription.
| Figure 3. Plasmid backbone carrying individual sgRNAs in the GeneCopoeia GeneHero™ CRISPR sgRNA libraries. |
Another significant advantage of the GeneCopoeia GeneHero™ CRISPR sgRNA libraries is that, because the sgRNA clones are individually constructed before pooling, researchers can order custom libraries in which we “mix and match” sgRNA clones from different libraries. In addition, we can also create custom CRISPR sgRNA libraries targeting new sets of genes.
Related products and services
In addition to our versatile, high-quality CRISPR sgRNA libraries, we also offer a number of companion products and services (Table 2). These include pre-made cell lines stably expressing Cas9 nuclease from the human AAVS1 Safe Harbor locus, and donor clones that you can use with our human AAVS1 Safe Harbor knockin kits to knock Cas9 in to AAVS1 in your cell line of choice. We also offer stable cell line services, in which we can construct a cell line carrying virtually any modification of interest, including expression of Cas9 from AAVS1 in a chosen cell line.
Conclusions
In this Technical Note, we highlighted a new and exciting technological development in the ever-evolving world of Genome Editing: CRISPR sgRNA libraries. As an alternative to whole genome libraries, GeneCopoeia provides several powerful, high-quality, and versatile CRISPR sgRNA libraries targeting smaller, focused pathways and gene groups. To find out more about how the GeneCopoeia GeneHero™ CRISPR sgRNA libraries can accelerate your research, visit our website: https://www.genecopoeia.com, or call us at 301-762-0888.
Custom stable cell line services available on request. Contact us for a quote.
Table 2. Companion products and services for the GeneCopoeia GeneHero™ CRISPR sgRNA libraries.
References
Cooper & Brockdorff (2013). Genome-wide shRNA screening to identify factors mediating Gata6 repression in mouse embryonic stem cells. Development 140, 4110.
Sethi, et al., (2012). An RNA interference lethality screen of the human druggable genome to identify molecular vulnerabilities in epithelial ovarian cancer. PLoSONE 7.
Shalem, et al. (2014). Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343, 84.
Sanjana, et al. (2014). Improved vectors and genome-wide libraries for CRISPR screening. Nature Methods 11, 783.
Wang, et al. (2014). Genetic screens in human cells using the CRISPR-Cas9 system. Science 343, 80.
Zhang, et al. (2014). Functional genomic screen of human stem cell differentiation reveals pathways involved in neurodevelopment and neurodegeneration. Proc Natl Acad Sci U S A 110, 12361.
| Copyright ©2020 GeneCopoeia, Inc. www.genecopoeia.com TNGE6-041715 |